| 产品: | 磷酸化 MYPT1 (Ser472) 抗体 |
| 货号: | AF3779 |
| 描述: | Rabbit polyclonal antibody to Phospho-MYPT1 (Ser472) |
| 应用: | WB |
| 文献验证: | WB |
| 反应: | Human, Mouse, Rat |
| 分子量: | 115kD(Calculated). |
| 蛋白号: | O14974 |
| RRID: | AB_2847093 |
产品描述
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: 适用于变性蛋白样本的免疫印迹检测. IHC: 适用于组织样本的石蜡(IHC-p)或冰冻(IHC-f)切片样本的免疫组化/荧光检测. IF/ICC: 适用于细胞样本的荧光检测. ELISA(peptide): 适用于抗原肽的ELISA检测.
引用格式: Affinity Biosciences Cat# AF3779, RRID:AB_2847093.
展开/折叠
M130; MBS; Myosin binding subunit; Myosin phosphatase target subunit 1; Myosin phosphatase targeting subunit 1; Myosin phosphatase-targeting subunit 1; MYPT1; MYPT1_HUMAN; PPP1R12A; Protein phosphatase 1 regulatory inhibitor subunit 12A; Protein phosphatase 1 regulatory subunit 12A; Protein phosphatase 1, regulatory (inhibitor) subunit 12A; Protein phosphatase myosin binding subunit; Protein phosphatase myosin-binding subunit;
抗原和靶标
A synthesized peptide derived from human MYPT1 around the phosphorylation site of Ser472.
Expressed in striated muscles, specifically in type 2a fibers (at protein level).
- O14974 MYPT1_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MKMADAKQKRNEQLKRWIGSETDLEPPVVKRQKTKVKFDDGAVFLAACSSGDTDEVLKLLHRGADINYANVDGLTALHQACIDDNVDMVKFLVENGANINQPDNEGWIPLHAAASCGYLDIAEFLIGQGAHVGAVNSEGDTPLDIAEEEAMEELLQNEVNRQGVDIEAARKEEERIMLRDARQWLNSGHINDVRHAKSGGTALHVAAAKGYTEVLKLLIQAGYDVNIKDYDGWTPLHAAAHWGKEEACRILVDNLCDMEMVNKVGQTAFDVADEDILGYLEELQKKQNLLHSEKRDKKSPLIESTANMDNNQSQKTFKNKETLIIEPEKNASRIESLEQEKVDEEEEGKKDESSCSSEEDEEDDSESEAETDKTKPLASVTNANTSSTQAAPVAVTTPTVSSGQATPTSPIKKFPTTATKISPKEEERKDESPATWRLGLRKTGSYGALAEITASKEGQKEKDTAGVTRSASSPRLSSSLDNKEKEKDSKGTRLAYVAPTIPRRLASTSDIEEKENRDSSSLRTSSSYTRRKWEDDLKKNSSVNEGSTYHKSCSFGRRQDDLISSSVPSTTSTPTVTSAAGLQKSLLSSTSTTTKITTGSSSAGTQSSTSNRLWAEDSTEKEKDSVPTAVTIPVAPTVVNAAASTTTLTTTTAGTVSSTTEVRERRRSYLTPVRDEESESQRKARSRQARQSRRSTQGVTLTDLQEAEKTIGRSRSTRTREQENEEKEKEEKEKQDKEKQEEKKESETSREDEYKQKYSRTYDETYQRYRPVSTSSSTTPSSSLSTMSSSLYASSQLNRPNSLVGITSAYSRGITKENEREGEKREEEKEGEDKSQPKSIRERRRPREKRRSTGVSFWTQDSDENEQEQQSDTEEGSNKKETQTDSISRYETSSTSAGDRYDSLLGRSGSYSYLEERKPYSSRLEKDDSTDFKKLYEQILAENEKLKAQLHDTNMELTDLKLQLEKATQRQERFADRSLLEMEKRERRALERRISEMEEELKMLPDLKADNQRLKDENGALIRVISKLSK
研究背景
Key regulator of protein phosphatase 1C (PPP1C). Mediates binding to myosin. As part of the PPP1C complex, involved in dephosphorylation of PLK1. Capable of inhibiting HIF1AN-dependent suppression of HIF1A activity.
Phosphorylated by CIT (Rho-associated kinase) (By similarity). Phosphorylated cooperatively by ROCK1 and CDC42BP on Thr-696. Phosphorylated on upon DNA damage, probably by ATM or ATR. In vitro, phosphorylation of Ser-695 by PKA and PKG appears to prevent phosphorylation of the inhibitory site Thr-696, probably mediated by PRKG1. Phosphorylation at Ser-445, Ser-472 and Ser-910 by NUAK1 promotes interaction with 14-3-3, leading to inhibit interaction with myosin light chain MLC2, preventing dephosphorylation of MLC2. May be phosphorylated at Thr-696 by DMPK; may inhibit the myosin phosphatase activity. Phosphorylated at Ser-473 by CDK1 during mitosis, creating docking sites for the POLO box domains of PLK1. Subsequently, PLK1 binds and phosphorylates PPP1R12A.
Cytoplasm. Cytoplasm>Cytoskeleton>Stress fiber.
Note: Also along actomyosin filaments.
Expressed in striated muscles, specifically in type 2a fibers (at protein level).
Heterotetramerization is mediated by the interaction between a coiled-coil of PRKG1 and the leucine/isoleucine zipper of PPP1R12A/MBS, the myosin-binding subunit of the myosin phosphatase.
The KVKF motif mediates interaction with PPP1CB.
研究领域
· Cellular Processes > Cellular community - eukaryotes > Focal adhesion. (View pathway)
· Cellular Processes > Cell motility > Regulation of actin cytoskeleton. (View pathway)
· Environmental Information Processing > Signal transduction > cGMP-PKG signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > cAMP signaling pathway. (View pathway)
· Human Diseases > Cancers: Overview > Proteoglycans in cancer.
· Organismal Systems > Circulatory system > Vascular smooth muscle contraction. (View pathway)
· Organismal Systems > Immune system > Platelet activation. (View pathway)
· Organismal Systems > Endocrine system > Oxytocin signaling pathway.
文献引用
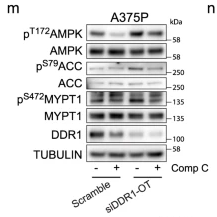
Application: WB Species: Human Sample: A375P cells
Application: WB Species: Human Sample: A375P cells
Application: WB Species: Mouse Sample: WM983B and A375M2 cells
限制条款
产品的规格、报价、验证数据请以官网为准,官网链接:www.affbiotech.com | www.affbiotech.cn(简体中文)| www.affbiotech.jp(日本語)产品的数据信息为Affinity所有,未经授权不得收集Affinity官网数据或资料用于商业用途,对抄袭产品数据的行为我们将保留诉诸法律的权利。
产品相关数据会因产品批次、产品检测情况随时调整,如您已订购该产品,请以订购时随货说明书为准,否则请以官网内容为准,官网内容有改动时恕不另行通知。
Affinity保证所销售产品均经过严格质量检测。如您购买的商品在规定时间内出现问题需要售后时,请您在Affinity官方渠道提交售后申请。产品仅供科学研究使用。不用于诊断和治疗。
产品未经授权不得转售。
Affinity Biosciences将不会对在使用我们的产品时可能发生的专利侵权或其他侵权行为负责。Affinity Biosciences, Affinity Biosciences标志和所有其他商标所有权归Affinity Biosciences LTD.