产品描述
*The optimal dilutions should be determined by the end user.
*Tips:
WB: 适用于变性蛋白样本的免疫印迹检测. IHC: 适用于组织样本的石蜡(IHC-p)或冰冻(IHC-f)切片样本的免疫组化/荧光检测. IF/ICC: 适用于细胞样本的荧光检测. ELISA(peptide): 适用于抗原肽的ELISA检测.
引用格式: Affinity Biosciences Cat# AF6126, RRID:AB_2835010.
展开/折叠
90 kda heat shock protein beta HSP90 beta; D6S182; FLJ26984; Heat shock 84 kDa; Heat shock 90kD protein 1, beta; Heat shock 90kDa protein 1 beta; Heat shock protein 90 alpha family class B member 1; Heat shock protein 90 kDa; Heat shock protein 90kDa alpha (cytosolic) class B member 1; Heat shock protein 90kDa alpha family class B member 1; Heat shock protein beta; Heat shock protein HSP 90 beta; Heat shock protein HSP 90-beta; HS90B_HUMAN; HSP 84; HSP 90; HSP 90 b; HSP 90b; HSP84; HSP90 BETA; hsp90ab1; HSP90B; HSPC2; HSPCB;
抗原和靶标
- P08238 HS90B_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MPEEVHHGEEEVETFAFQAEIAQLMSLIINTFYSNKEIFLRELISNASDALDKIRYESLTDPSKLDSGKELKIDIIPNPQERTLTLVDTGIGMTKADLINNLGTIAKSGTKAFMEALQAGADISMIGQFGVGFYSAYLVAEKVVVITKHNDDEQYAWESSAGGSFTVRADHGEPIGRGTKVILHLKEDQTEYLEERRVKEVVKKHSQFIGYPITLYLEKEREKEISDDEAEEEKGEKEEEDKDDEEKPKIEDVGSDEEDDSGKDKKKKTKKIKEKYIDQEELNKTKPIWTRNPDDITQEEYGEFYKSLTNDWEDHLAVKHFSVEGQLEFRALLFIPRRAPFDLFENKKKKNNIKLYVRRVFIMDSCDELIPEYLNFIRGVVDSEDLPLNISREMLQQSKILKVIRKNIVKKCLELFSELAEDKENYKKFYEAFSKNLKLGIHEDSTNRRRLSELLRYHTSQSGDEMTSLSEYVSRMKETQKSIYYITGESKEQVANSAFVERVRKRGFEVVYMTEPIDEYCVQQLKEFDGKSLVSVTKEGLELPEDEEEKKKMEESKAKFENLCKLMKEILDKKVEKVTISNRLVSSPCCIVTSTYGWTANMERIMKAQALRDNSTMGYMMAKKHLEINPDHPIVETLRQKAEADKNDKAVKDLVVLLFETALLSSGFSLEDPQTHSNRIYRMIKLGLGIDEDEVAAEEPNAAVPDEIPPLEGDEDASRMEEVD
种属预测
score>80的预测可信度较高,可尝试用于WB检测。*预测模型主要基于免疫原序列比对,结果仅作参考,不作为质保凭据。
High(score>80) Medium(80>score>50) Low(score<50) No confidence
研究背景
Molecular chaperone that promotes the maturation, structural maintenance and proper regulation of specific target proteins involved for instance in cell cycle control and signal transduction. Undergoes a functional cycle that is linked to its ATPase activity. This cycle probably induces conformational changes in the client proteins, thereby causing their activation. Interacts dynamically with various co-chaperones that modulate its substrate recognition, ATPase cycle and chaperone function. Engages with a range of client protein classes via its interaction with various co-chaperone proteins or complexes, that act as adapters, simultaneously able to interact with the specific client and the central chaperone itself. Recruitment of ATP and co-chaperone followed by client protein forms a functional chaperone. After the completion of the chaperoning process, properly folded client protein and co-chaperone leave HSP90 in an ADP-bound partially open conformation and finally, ADP is released from HSP90 which acquires an open conformation for the next cycle. Apart from its chaperone activity, it also plays a role in the regulation of the transcription machinery. HSP90 and its co-chaperones modulate transcription at least at three different levels. In the first place, they alter the steady-state levels of certain transcription factors in response to various physiological cues. Second, they modulate the activity of certain epigenetic modifiers, such as histone deacetylases or DNA methyl transferases, and thereby respond to the change in the environment. Third, they participate in the eviction of histones from the promoter region of certain genes and thereby turn on gene expression. Antagonizes STUB1-mediated inhibition of TGF-beta signaling via inhibition of STUB1-mediated SMAD3 ubiquitination and degradation. Promotes cell differentiation by chaperoning BIRC2 and thereby protecting from auto-ubiquitination and degradation by the proteasomal machinery. Main chaperone that is involved in the phosphorylation/activation of the STAT1 by chaperoning both JAK2 and PRKCE under heat shock and in turn, activates its own transcription.
Ubiquitinated in the presence of STUB1-UBE2D1 complex (in vitro).
ISGylated.
S-nitrosylated; negatively regulates the ATPase activity.
Phosphorylation at Tyr-301 by SRC is induced by lipopolysaccharide. Phosphorylation at Ser-226 and Ser-255 inhibits AHR interaction.
Methylated by SMYD2; facilitates dimerization and chaperone complex formation; promotes cancer cell proliferation.
Cleaved following oxidative stress resulting in HSP90AB1 protein radicals formation; disrupts the chaperoning function and the degradation of its client proteins.
Cytoplasm. Melanosome. Nucleus. Secreted. Cell membrane.
Note: Identified by mass spectrometry in melanosome fractions from stage I to stage IV (PubMed:17081065). Translocates with BIRC2 from the nucleus to the cytoplasm during differentiation (PubMed:18239673). Secreted when associated with TGFB1 processed form (LAP) (PubMed:20599762).
Monomer. Homodimer. Forms a complex with CDK6 and CDC37. Interacts with UNC45A; binding to UNC45A involves 2 UNC45A monomers per HSP90AB1 dimer. Interacts with CHORDC1 (By similarity). Interacts with DNAJC7. Interacts with FKBP4. May interact with NWD1. Interacts with SGTA. Interacts with HSF1 in an ATP-dependent manner. Interacts with MET; the interaction suppresses MET kinase activity. Interacts with ERBB2 in an ATP-dependent manner; the interaction suppresses ERBB2 kinase activity. Interacts with HIF1A, KEAP1 and RHOBTB2. Interacts with STUB1 and SMAD3. Interacts with XPO1 and AHSA1. Interacts with BIRC2. Interacts with KCNQ4; promotes cell surface expression of KCNQ4. Interacts with BIRC2; prevents auto-ubiquitination and degradation of its client protein BIRC2. Interacts with NOS3. Interacts with AHR; interaction is inhibited by HSP90AB1 phosphorylation on Ser-226 and Ser-255. Interacts with STIP1 and CDC37; upon SMYD2-dependent methylation. Interacts with JAK2 and PRKCE; promotes functional activation in a heat shock-dependent manner. Interacts with HSP90AA1; interaction is constitutive. HSP90AB1-CDC37 chaperone complex interacts with inactive MAPK7 (via N-terminal half) in resting cells; the interaction is MAP2K5-independent and prevents from ubiquitination and proteasomal degradation. Interacts with CDC25A; prevents heat shock-mediated CDC25A degradation and contributes to cell cycle progression. Interacts with TP53 (via DNA binding domain); suppresses TP53 aggregation and prevents from irreversible thermal inactivation. Interacts with TGFB1 processed form (LAP); inhibits latent TGFB1 activation. Interacts with TRIM8; prevents nucleus translocation of phosphorylated STAT3 and HSP90AB1 (By similarity). Interacts with NR3C1 (via domain NR LBD) and NR1D1 (via domain NR LBD) (By similarity). Interacts with PDCL3 (By similarity). Interacts with TTC4 (via TPR repeats).
The TPR repeat-binding motif mediates interaction with TPR repeat-containing proteins.
Belongs to the heat shock protein 90 family.
研究领域
· Cellular Processes > Cell growth and death > Necroptosis. (View pathway)
· Environmental Information Processing > Signal transduction > PI3K-Akt signaling pathway. (View pathway)
· Genetic Information Processing > Folding, sorting and degradation > Protein processing in endoplasmic reticulum. (View pathway)
· Human Diseases > Cancers: Overview > Pathways in cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Prostate cancer. (View pathway)
· Organismal Systems > Immune system > Antigen processing and presentation. (View pathway)
· Organismal Systems > Immune system > NOD-like receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > IL-17 signaling pathway. (View pathway)
· Organismal Systems > Immune system > Th17 cell differentiation. (View pathway)
· Organismal Systems > Endocrine system > Progesterone-mediated oocyte maturation.
· Organismal Systems > Endocrine system > Estrogen signaling pathway. (View pathway)
文献引用
Application: WB Species: human Sample: A549 cells
Application: WB Species: human Sample: A549 cells
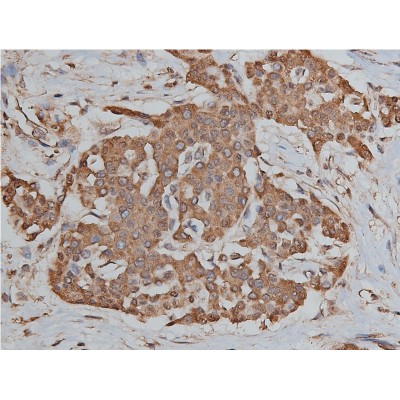
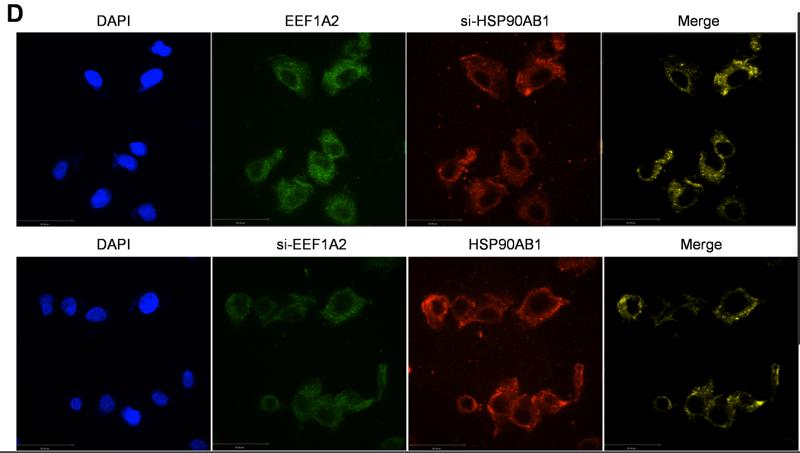
Application: IF/ICC Species: Human Sample: ESCC
Application: WB Species: Human Sample: ESCC
限制条款
产品的规格、报价、验证数据请以官网为准,官网链接:www.affbiotech.com | www.affbiotech.cn(简体中文)| www.affbiotech.jp(日本語)产品的数据信息为Affinity所有,未经授权不得收集Affinity官网数据或资料用于商业用途,对抄袭产品数据的行为我们将保留诉诸法律的权利。
产品相关数据会因产品批次、产品检测情况随时调整,如您已订购该产品,请以订购时随货说明书为准,否则请以官网内容为准,官网内容有改动时恕不另行通知。
Affinity保证所销售产品均经过严格质量检测。如您购买的商品在规定时间内出现问题需要售后时,请您在Affinity官方渠道提交售后申请。产品仅供科学研究使用。不用于诊断和治疗。
产品未经授权不得转售。
Affinity Biosciences将不会对在使用我们的产品时可能发生的专利侵权或其他侵权行为负责。Affinity Biosciences, Affinity Biosciences标志和所有其他商标所有权归Affinity Biosciences LTD.