产品描述
*The optimal dilutions should be determined by the end user.
*Tips:
WB: 适用于变性蛋白样本的免疫印迹检测. IHC: 适用于组织样本的石蜡(IHC-p)或冰冻(IHC-f)切片样本的免疫组化/荧光检测. IF/ICC: 适用于细胞样本的荧光检测. ELISA(peptide): 适用于抗原肽的ELISA检测.
引用格式: Affinity Biosciences Cat# AF6319, RRID:AB_2835178.
展开/折叠
C Jun kinase 2; c Jun N terminal kinase 1; c Jun N terminal kinase 2; c Jun N terminal kinase 3; c-Jun N-terminal kinase 1; JNK 46; JNK 55; JNK; JNK-46; JNK1; JNK1A2; JNK2; JNK21B1/2; JNK2A; JNK2ALPHA; JNK2B; JNK2BETA; JNK3 alpha protein kinase; JNK3; JNK3A; Jun kinase; JUN N terminal kinase; MAP kinase 10; MAP kinase 8; MAP kinase 9; MAP kinase p49 3F12; MAPK 10; MAPK 8; MAPK 9; MAPK10; mapk8; MAPK9; Mitogen activated protein kinase 10; Mitogen activated protein kinase 8; Mitogen activated protein kinase 8 isoform JNK1 alpha1; Mitogen activated protein kinase 8 isoform JNK1 beta2; Mitogen activated protein kinase 9; Mitogen-activated protein kinase 8; MK08_HUMAN; p493F12; p54a; p54aSAPK; p54bSAPK; PRKM10; PRKM8; PRKM9; SAPK; SAPK(beta); SAPK1; SAPK1a; SAPK1b; SAPK1c; Stress activated protein kinase 1; Stress activated protein kinase 1a; Stress activated protein kinase 1b; Stress activated protein kinase 1c; Stress activated protein kinase beta; Stress activated protein kinase JNK1; Stress activated protein kinase JNK2; Stress activated protein kinase JNK3; Stress-activated protein kinase 1c; Stress-activated protein kinase JNK1; c Jun kinase 2; C Jun N terminal kinase 2; c-Jun N-terminal kinase 2; JNK 55; JNK-55; JNK2 alpha; JNK2; JNK2 beta; JNK2A; JNK2alpha; JNK2B; JNK2BETA; Jun kinase; MAP kinase 9; MAPK 9; Mapk9; Mitogen activated protein kinase 9; Mitogen-activated protein kinase 9; MK09_HUMAN; P54a; p54aSAPK; PRKM9; Protein kinase, mitogen-activated, 9; SAPK alpha; SAPK; SAPK1a; Stress activated protein kinase 1a; Stress-activated protein kinase JNK2; c Jun kinase 3; c-Jun N-terminal kinase 3; cJun N terminal kinase 3; FLJ12099; FLJ33785; JNK3 alpha protein kinase; JNK3; JNK3A; MAP kinase 10; MAP kinase; MAP kinase p49 3F12; MAPK 10; Mapk10; MGC50974; mitogen activated protein kinase 10; Mitogen-activated protein kinase 10; MK10_HUMAN; p493F12; p54bSAPK; PRKM10; protein kinase mitogen activated 10; SAPK1b; Stress activated protein kinase 1b; stress activated protein kinase beta; Stress activated protein kinase JNK3; Stress-activated protein kinase JNK3;
抗原和靶标
Specific to a subset of neurons in the nervous system. Present in the hippocampus and areas, cerebellum, striatum, brain stem, and weakly in the spinal cord. Very weak expression in testis and kidney.
- P45983 MK08_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MSRSKRDNNFYSVEIGDSTFTVLKRYQNLKPIGSGAQGIVCAAYDAILERNVAIKKLSRPFQNQTHAKRAYRELVLMKCVNHKNIIGLLNVFTPQKSLEEFQDVYIVMELMDANLCQVIQMELDHERMSYLLYQMLCGIKHLHSAGIIHRDLKPSNIVVKSDCTLKILDFGLARTAGTSFMMTPYVVTRYYRAPEVILGMGYKENVDLWSVGCIMGEMVCHKILFPGRDYIDQWNKVIEQLGTPCPEFMKKLQPTVRTYVENRPKYAGYSFEKLFPDVLFPADSEHNKLKASQARDLLSKMLVIDASKRISVDEALQHPYINVWYDPSEAEAPPPKIPDKQLDEREHTIEEWKELIYKEVMDLEERTKNGVIRGQPSPLGAAVINGSQHPSSSSSVNDVSSMSTDPTLASDTDSSLEAAAGPLGCCR
- P45984 MK09_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MSDSKCDSQFYSVQVADSTFTVLKRYQQLKPIGSGAQGIVCAAFDTVLGINVAVKKLSRPFQNQTHAKRAYRELVLLKCVNHKNIISLLNVFTPQKTLEEFQDVYLVMELMDANLCQVIHMELDHERMSYLLYQMLCGIKHLHSAGIIHRDLKPSNIVVKSDCTLKILDFGLARTACTNFMMTPYVVTRYYRAPEVILGMGYKENVDIWSVGCIMGELVKGCVIFQGTDHIDQWNKVIEQLGTPSAEFMKKLQPTVRNYVENRPKYPGIKFEELFPDWIFPSESERDKIKTSQARDLLSKMLVIDPDKRISVDEALRHPYITVWYDPAEAEAPPPQIYDAQLEEREHAIEEWKELIYKEVMDWEERSKNGVVKDQPSDAAVSSNATPSQSSSINDISSMSTEQTLASDTDSSLDASTGPLEGCR
- P53779 MK10_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MSLHFLYYCSEPTLDVKIAFCQGFDKQVDVSYIAKHYNMSKSKVDNQFYSVEVGDSTFTVLKRYQNLKPIGSGAQGIVCAAYDAVLDRNVAIKKLSRPFQNQTHAKRAYRELVLMKCVNHKNIISLLNVFTPQKTLEEFQDVYLVMELMDANLCQVIQMELDHERMSYLLYQMLCGIKHLHSAGIIHRDLKPSNIVVKSDCTLKILDFGLARTAGTSFMMTPYVVTRYYRAPEVILGMGYKENVDIWSVGCIMGEMVRHKILFPGRDYIDQWNKVIEQLGTPCPEFMKKLQPTVRNYVENRPKYAGLTFPKLFPDSLFPADSEHNKLKASQARDLLSKMLVIDPAKRISVDDALQHPYINVWYDPAEVEAPPPQIYDKQLDEREHTIEEWKELIYKEVMNSEEKTKNGVVKGQPSPSGAAVNSSESLPPSSSVNDISSMSTDQTLASDTDSSLEASAGPLGCCR
种属预测
score>80的预测可信度较高,可尝试用于WB检测。*预测模型主要基于免疫原序列比对,结果仅作参考,不作为质保凭据。
High(score>80) Medium(80>score>50) Low(score<50) No confidence
研究背景
Serine/threonine-protein kinase involved in various processes such as cell proliferation, differentiation, migration, transformation and programmed cell death. Extracellular stimuli such as proinflammatory cytokines or physical stress stimulate the stress-activated protein kinase/c-Jun N-terminal kinase (SAP/JNK) signaling pathway. In this cascade, two dual specificity kinases MAP2K4/MKK4 and MAP2K7/MKK7 phosphorylate and activate MAPK8/JNK1. In turn, MAPK8/JNK1 phosphorylates a number of transcription factors, primarily components of AP-1 such as JUN, JDP2 and ATF2 and thus regulates AP-1 transcriptional activity. Phosphorylates the replication licensing factor CDT1, inhibiting the interaction between CDT1 and the histone H4 acetylase HBO1 to replication origins. Loss of this interaction abrogates the acetylation required for replication initiation. Promotes stressed cell apoptosis by phosphorylating key regulatory factors including p53/TP53 and Yes-associates protein YAP1. In T-cells, MAPK8 and MAPK9 are required for polarized differentiation of T-helper cells into Th1 cells. Contributes to the survival of erythroid cells by phosphorylating the antagonist of cell death BAD upon EPO stimulation. Mediates starvation-induced BCL2 phosphorylation, BCL2 dissociation from BECN1, and thus activation of autophagy. Phosphorylates STMN2 and hence regulates microtubule dynamics, controlling neurite elongation in cortical neurons. In the developing brain, through its cytoplasmic activity on STMN2, negatively regulates the rate of exit from multipolar stage and of radial migration from the ventricular zone. Phosphorylates several other substrates including heat shock factor protein 4 (HSF4), the deacetylase SIRT1, ELK1, or the E3 ligase ITCH. Phosphorylates the CLOCK-ARNTL/BMAL1 heterodimer and plays a role in the regulation of the circadian clock. Phosphorylates the heat shock transcription factor HSF1, suppressing HSF1-induced transcriptional activity. Phosphorylates POU5F1, which results in the inhibition of POU5F1's transcriptional activity and enhances its proteosomal degradation (By similarity). Phosphorylates JUND and this phosphorylation is inhibited in the presence of MEN1 (By similarity). In neurons, phosphorylates SYT4 which captures neuronal dense core vesicles at synapses (By similarity).
JNK1 isoforms display different binding patterns: beta-1 preferentially binds to c-Jun, whereas alpha-1, alpha-2, and beta-2 have a similar low level of binding to both c-Jun or ATF2. However, there is no correlation between binding and phosphorylation, which is achieved at about the same efficiency by all isoforms.
Dually phosphorylated on Thr-183 and Tyr-185 by MAP2K7 and MAP2K4, which activates the enzyme. Phosphorylated by TAOK2. May be phosphorylated at Thr-183 and Tyr-185 by MAP3K1/MEKK1. Phosphorylated form is more concentrated at synapses than none-phosphorylated (By similarity).
Cytoplasm. Nucleus. Cell junction>Synapse.
Note: In the cortical neurons, predominantly cytoplasmic and associated with the Golgi apparatus and endosomal fraction. Increased neuronal activity increases phosphorylated form at synapses (By similarity). Colocalizes with POU5F1 in the nucleus.
Binds to at least four scaffolding proteins, MAPK8IP1/JIP-1, MAPK8IP2/JIP-2, MAPK8IP3/JIP-3/JSAP1 and SPAG9/MAPK8IP4/JIP-4. These proteins also bind other components of the JNK signaling pathway. Interacts with TP53 and WWOX. Interacts with JAMP (By similarity). Forms a complex with MAPK8IP1 and ARHGEF28 (By similarity). Interacts with HSF1 (via D domain and preferentially with hyperphosphorylated form); this interaction occurs under both normal growth conditions and immediately upon heat shock. Interacts (phosphorylated form) with NFE2; the interaction phosphorylates NFE2 in undifferentiated cells (By similarity). Interacts with NFATC4. Interacts with MECOM; regulates JNK signaling. Interacts with PIN1; this interaction mediates MAPK8 conformational changes leading to the binding of MAPK8 to its substrates. Interacts with GRIPAP1. Interacts with POU5F1; phosphorylates POU5F1 at 'Ser-355'. Found in a complex with SH3RF1, RAC1, MAP3K11/MLK3, MAP2K7/MKK7 and MAPK8IP1/JIP1. Found in a complex with SH3RF1, RAC2, MAP3K7/TAK1, MAP2K7/MKK7, MAPK8IP1/JIP1 and MAPK9/JNK2 (By similarity).
The TXY motif contains the threonine and tyrosine residues whose phosphorylation activates the MAP kinases.
Belongs to the protein kinase superfamily. CMGC Ser/Thr protein kinase family. MAP kinase subfamily.
Serine/threonine-protein kinase involved in various processes such as cell proliferation, differentiation, migration, transformation and programmed cell death. Extracellular stimuli such as proinflammatory cytokines or physical stress stimulate the stress-activated protein kinase/c-Jun N-terminal kinase (SAP/JNK) signaling pathway. In this cascade, two dual specificity kinases MAP2K4/MKK4 and MAP2K7/MKK7 phosphorylate and activate MAPK9/JNK2. In turn, MAPK9/JNK2 phosphorylates a number of transcription factors, primarily components of AP-1 such as JUN and ATF2 and thus regulates AP-1 transcriptional activity. In response to oxidative or ribotoxic stresses, inhibits rRNA synthesis by phosphorylating and inactivating the RNA polymerase 1-specific transcription initiation factor RRN3. Promotes stressed cell apoptosis by phosphorylating key regulatory factors including TP53 and YAP1. In T-cells, MAPK8 and MAPK9 are required for polarized differentiation of T-helper cells into Th1 cells. Upon T-cell receptor (TCR) stimulation, is activated by CARMA1, BCL10, MAP2K7 and MAP3K7/TAK1 to regulate JUN protein levels. Plays an important role in the osmotic stress-induced epithelial tight-junctions disruption. When activated, promotes beta-catenin/CTNNB1 degradation and inhibits the canonical Wnt signaling pathway. Participates also in neurite growth in spiral ganglion neurons. Phosphorylates the CLOCK-ARNTL/BMAL1 heterodimer and plays a role in the regulation of the circadian clock. Phosphorylates POU5F1, which results in the inhibition of POU5F1's transcriptional activity and enhances its proteosomal degradation (By similarity).
MAPK9 isoforms display different binding patterns: alpha-1 and alpha-2 preferentially bind to JUN, whereas beta-1 and beta-2 bind to ATF2. However, there is no correlation between binding and phosphorylation, which is achieved at about the same efficiency by all isoforms. JUNB is not a substrate for JNK2 alpha-2, and JUND binds only weakly to it.
Dually phosphorylated on Thr-183 and Tyr-185 by MAP2K7 and MAP2K4, which activates the enzyme. Autophosphorylated in vitro.
Cytoplasm. Nucleus.
Note: Colocalizes with POU5F1 in the nucleus.
Interacts with MECOM (By similarity). Interacts with DCLK2 (By similarity). Binds to at least four scaffolding proteins, MAPK8IP1/JIP-1, MAPK8IP2/JIP-2, MAPK8IP3/JIP-3/JSAP1 and SPAG9/MAPK8IP4/JIP-4. These proteins also bind other components of the JNK signaling pathway (By similarity). Interacts with NFATC4. Interacts with ATF7; the interaction does not phosphorylate ATF7 but acts as a docking site for ATF7-associated partners such as JUN. Interacts with BCL10. Interacts with CTNNB1 and GSK3B. Interacts with MAPKBP1. Interacts with POU5F1; phosphorylates POU5F1 at 'Ser-355'. Found in a complex with SH3RF1, RAC2, MAP3K7/TAK1, MAP2K7/MKK7, MAPK8IP1/JIP1 and MAPK8/JNK1 (By similarity).
The TXY motif contains the threonine and tyrosine residues whose phosphorylation activates the MAP kinases.
Belongs to the protein kinase superfamily. CMGC Ser/Thr protein kinase family. MAP kinase subfamily.
Serine/threonine-protein kinase involved in various processes such as neuronal proliferation, differentiation, migration and programmed cell death. Extracellular stimuli such as proinflammatory cytokines or physical stress stimulate the stress-activated protein kinase/c-Jun N-terminal kinase (SAP/JNK) signaling pathway. In this cascade, two dual specificity kinases MAP2K4/MKK4 and MAP2K7/MKK7 phosphorylate and activate MAPK10/JNK3. In turn, MAPK10/JNK3 phosphorylates a number of transcription factors, primarily components of AP-1 such as JUN and ATF2 and thus regulates AP-1 transcriptional activity. Plays regulatory roles in the signaling pathways during neuronal apoptosis. Phosphorylates the neuronal microtubule regulator STMN2. Acts in the regulation of the amyloid-beta precursor protein/APP signaling during neuronal differentiation by phosphorylating APP. Participates also in neurite growth in spiral ganglion neurons. Phosphorylates the CLOCK-ARNTL/BMAL1 heterodimer and plays a role in the photic regulation of the circadian clock. Phosphorylates JUND and this phosphorylation is inhibited in the presence of MEN1.
Dually phosphorylated on Thr-221 and Tyr-223 by MAP2K4 and MAP2K7, which activates the enzyme. MAP2K7 shows a strong preference for Thr-221 while MAP2K4 phosphorylates Tyr-223 preferentially. Weakly autophosphorylated on threonine and tyrosine residues in vitro.
Palmitoylation regulates subcellular location and axonal development.
Cytoplasm. Membrane>Lipid-anchor. Nucleus. Mitochondrion.
Note: Palmitoylation regulates MAPK10 trafficking to cytoskeleton. Recruited to the mitochondria in the presence of SARM1 (By similarity).
Specific to a subset of neurons in the nervous system. Present in the hippocampus and areas, cerebellum, striatum, brain stem, and weakly in the spinal cord. Very weak expression in testis and kidney.
Interacts with MAPKBP1 (By similarity). Interacts with MAPK8IP1/JIP-1 and MAPK8IP3/JIP-3/JSAP1 (By similarity). Interacts with SPAG9/MAPK8IP4/JIP4. Interacts with HDAC9. Interacts with ARRB2; the interaction enhances MAPK10 activation by MAP3K5. Interacts with SARM1 (By similarity). Interacts with JUND; interaction is inhibited in the presence of MEN1.
The TXY motif contains the threonine and tyrosine residues whose phosphorylation activates the MAP kinases.
Belongs to the protein kinase superfamily. CMGC Ser/Thr protein kinase family. MAP kinase subfamily.
研究领域
· Cellular Processes > Transport and catabolism > Autophagy - animal. (View pathway)
· Cellular Processes > Cell growth and death > Apoptosis. (View pathway)
· Cellular Processes > Cell growth and death > Apoptosis - multiple species. (View pathway)
· Cellular Processes > Cell growth and death > Necroptosis. (View pathway)
· Cellular Processes > Cellular community - eukaryotes > Focal adhesion. (View pathway)
· Cellular Processes > Cellular community - eukaryotes > Tight junction. (View pathway)
· Environmental Information Processing > Signal transduction > MAPK signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > ErbB signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Ras signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > cAMP signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > FoxO signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Sphingolipid signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Wnt signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > TNF signaling pathway. (View pathway)
· Genetic Information Processing > Folding, sorting and degradation > Protein processing in endoplasmic reticulum. (View pathway)
· Human Diseases > Drug resistance: Antineoplastic > Endocrine resistance.
· Human Diseases > Endocrine and metabolic diseases > Type II diabetes mellitus.
· Human Diseases > Endocrine and metabolic diseases > Insulin resistance.
· Human Diseases > Endocrine and metabolic diseases > Non-alcoholic fatty liver disease (NAFLD).
· Human Diseases > Infectious diseases: Bacterial > Epithelial cell signaling in Helicobacter pylori infection.
· Human Diseases > Infectious diseases: Bacterial > Shigellosis.
· Human Diseases > Infectious diseases: Bacterial > Salmonella infection.
· Human Diseases > Infectious diseases: Bacterial > Pertussis.
· Human Diseases > Infectious diseases: Parasitic > Chagas disease (American trypanosomiasis).
· Human Diseases > Infectious diseases: Parasitic > Toxoplasmosis.
· Human Diseases > Infectious diseases: Bacterial > Tuberculosis.
· Human Diseases > Infectious diseases: Viral > Hepatitis C.
· Human Diseases > Infectious diseases: Viral > Hepatitis B.
· Human Diseases > Infectious diseases: Viral > Influenza A.
· Human Diseases > Infectious diseases: Viral > HTLV-I infection.
· Human Diseases > Infectious diseases: Viral > Herpes simplex infection.
· Human Diseases > Infectious diseases: Viral > Epstein-Barr virus infection.
· Human Diseases > Cancers: Overview > Pathways in cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Colorectal cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Pancreatic cancer. (View pathway)
· Human Diseases > Cancers: Overview > Choline metabolism in cancer. (View pathway)
· Organismal Systems > Development > Osteoclast differentiation. (View pathway)
· Organismal Systems > Immune system > Toll-like receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > NOD-like receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > RIG-I-like receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > IL-17 signaling pathway. (View pathway)
· Organismal Systems > Immune system > Th1 and Th2 cell differentiation. (View pathway)
· Organismal Systems > Immune system > Th17 cell differentiation. (View pathway)
· Organismal Systems > Immune system > T cell receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > Fc epsilon RI signaling pathway. (View pathway)
· Organismal Systems > Nervous system > Neurotrophin signaling pathway. (View pathway)
· Organismal Systems > Nervous system > Retrograde endocannabinoid signaling. (View pathway)
· Organismal Systems > Nervous system > Dopaminergic synapse.
· Organismal Systems > Sensory system > Inflammatory mediator regulation of TRP channels. (View pathway)
· Organismal Systems > Endocrine system > Insulin signaling pathway. (View pathway)
· Organismal Systems > Endocrine system > Progesterone-mediated oocyte maturation.
· Organismal Systems > Endocrine system > Prolactin signaling pathway. (View pathway)
· Organismal Systems > Endocrine system > Adipocytokine signaling pathway.
· Organismal Systems > Endocrine system > Relaxin signaling pathway.
文献引用
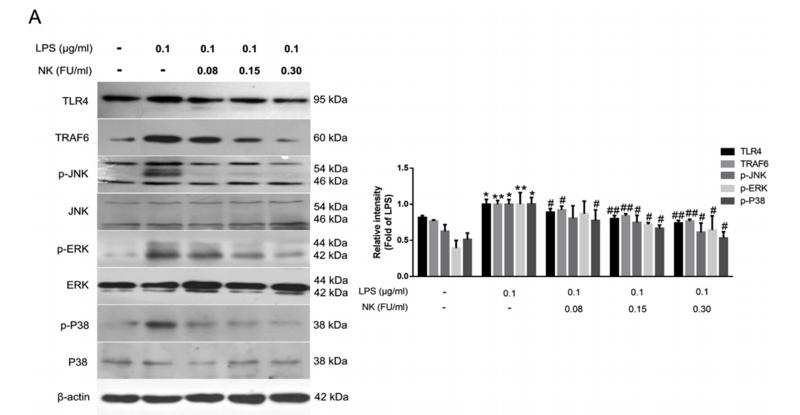
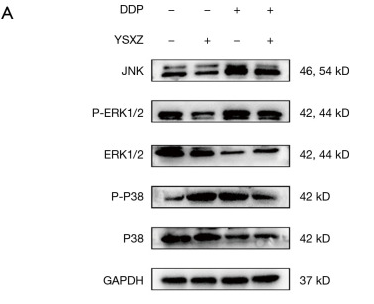
Application: WB Species: human Sample: HUVECs cells
限制条款
产品的规格、报价、验证数据请以官网为准,官网链接:www.affbiotech.com | www.affbiotech.cn(简体中文)| www.affbiotech.jp(日本語)产品的数据信息为Affinity所有,未经授权不得收集Affinity官网数据或资料用于商业用途,对抄袭产品数据的行为我们将保留诉诸法律的权利。
产品相关数据会因产品批次、产品检测情况随时调整,如您已订购该产品,请以订购时随货说明书为准,否则请以官网内容为准,官网内容有改动时恕不另行通知。
Affinity保证所销售产品均经过严格质量检测。如您购买的商品在规定时间内出现问题需要售后时,请您在Affinity官方渠道提交售后申请。产品仅供科学研究使用。不用于诊断和治疗。
产品未经授权不得转售。
Affinity Biosciences将不会对在使用我们的产品时可能发生的专利侵权或其他侵权行为负责。Affinity Biosciences, Affinity Biosciences标志和所有其他商标所有权归Affinity Biosciences LTD.