产品描述
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: 适用于变性蛋白样本的免疫印迹检测. IHC: 适用于组织样本的石蜡(IHC-p)或冰冻(IHC-f)切片样本的免疫组化/荧光检测. IF/ICC: 适用于细胞样本的荧光检测. ELISA(peptide): 适用于抗原肽的ELISA检测.
引用格式: Affinity Biosciences Cat# AF5376, RRID:AB_2810280.
展开/折叠
E3 ubiquitin-protein ligase TRAF6; Interleukin 1 signal transducer; Interleukin-1 signal transducer; MGC 3310; MGC:3310; MGC3310; OTTHUMP00000232772; OTTHUMP00000232773; RING finger protein 85; RNF 85; RNF85; TNF receptor associated factor 6; TNF receptor-associated factor 6; TNF receptor-associated factor 6, E3 ubiquitin protein ligase; TRAF 6; Traf6; TRAF6_HUMAN;
抗原和靶标
A synthesized peptide derived from human TRAF6, corresponding to a region within the internal amino acids.
Expressed in heart, brain, placenta, lung, liver, skeletal muscle, kidney and pancreas.
- Q9Y4K3 TRAF6_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MSLLNCENSCGSSQSESDCCVAMASSCSAVTKDDSVGGTASTGNLSSSFMEEIQGYDVEFDPPLESKYECPICLMALREAVQTPCGHRFCKACIIKSIRDAGHKCPVDNEILLENQLFPDNFAKREILSLMVKCPNEGCLHKMELRHLEDHQAHCEFALMDCPQCQRPFQKFHINIHILKDCPRRQVSCDNCAASMAFEDKEIHDQNCPLANVICEYCNTILIREQMPNHYDLDCPTAPIPCTFSTFGCHEKMQRNHLARHLQENTQSHMRMLAQAVHSLSVIPDSGYISEVRNFQETIHQLEGRLVRQDHQIRELTAKMETQSMYVSELKRTIRTLEDKVAEIEAQQCNGIYIWKIGNFGMHLKCQEEEKPVVIHSPGFYTGKPGYKLCMRLHLQLPTAQRCANYISLFVHTMQGEYDSHLPWPFQGTIRLTILDQSEAPVRQNHEEIMDAKPELLAFQRPTIPRNPKGFGYVTFMHLEALRQRTFIKDDTLLVRCEVSTRFDMGSLRREGFQPRSTDAGV
种属预测
score>80的预测可信度较高,可尝试用于WB检测。*预测模型主要基于免疫原序列比对,结果仅作参考,不作为质保凭据。
High(score>80) Medium(80>score>50) Low(score<50) No confidence
研究背景
E3 ubiquitin ligase that, together with UBE2N and UBE2V1, mediates the synthesis of 'Lys-63'-linked-polyubiquitin chains conjugated to proteins, such as IKBKG, IRAK1, AKT1 and AKT2. Also mediates ubiquitination of free/unanchored polyubiquitin chain that leads to MAP3K7 activation. Leads to the activation of NF-kappa-B and JUN. May be essential for the formation of functional osteoclasts. Seems to also play a role in dendritic cells (DCs) maturation and/or activation. Represses c-Myb-mediated transactivation, in B-lymphocytes. Adapter protein that seems to play a role in signal transduction initiated via TNF receptor, IL-1 receptor and IL-17 receptor. Regulates osteoclast differentiation by mediating the activation of adapter protein complex 1 (AP-1) and NF-kappa-B, in response to RANK-L stimulation. Together with MAP3K8, mediates CD40 signals that activate ERK in B-cells and macrophages, and thus may play a role in the regulation of immunoglobulin production.
Sumoylated on Lys-124, Lys-142 and Lys-453 with SUMO1.
Polyubiquitinated on Lys-124; after cell stimulation with IL-1-beta or TGF-beta. This ligand-induced cell stimulation leads to dimerization/oligomerization of TRAF6 molecules, followed by auto-ubiquitination which involves UBE2N and UBE2V1 and leads to TRAF6 activation. This 'Lys-63' site-specific poly-ubiquitination appears to be associated with the activation of signaling molecules. Endogenous autoubiquitination occurs only for the cytoplasmic form. Deubiquitinated by USP10 in a TANK-dependent manner, leading to the negative regulation of NF-kappaB signaling upon DNA damage.
Cytoplasm. Cytoplasm>Cell cortex. Nucleus. Lipid droplet.
Note: Found in the nuclei of some aggressive B-cell lymphoma cell lines as well as in the nuclei of both resting and activated T- and B-lymphocytes. Found in punctate nuclear body protein complexes. Ubiquitination may occur in the cytoplasm and sumoylation in the nucleus. RSAD2/viperin recruits it to the lipid droplet (By similarity).
Expressed in heart, brain, placenta, lung, liver, skeletal muscle, kidney and pancreas.
The coiled coil domain mediates homo- and hetero-oligomerization.
The MATH/TRAF domain binds to receptor cytoplasmic domains.
Belongs to the TNF receptor-associated factor family. A subfamily.
研究领域
· Cellular Processes > Transport and catabolism > Autophagy - animal. (View pathway)
· Cellular Processes > Transport and catabolism > Endocytosis. (View pathway)
· Environmental Information Processing > Signal transduction > MAPK signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > NF-kappa B signaling pathway. (View pathway)
· Genetic Information Processing > Folding, sorting and degradation > Ubiquitin mediated proteolysis. (View pathway)
· Human Diseases > Infectious diseases: Bacterial > Pertussis.
· Human Diseases > Infectious diseases: Parasitic > Leishmaniasis.
· Human Diseases > Infectious diseases: Parasitic > Chagas disease (American trypanosomiasis).
· Human Diseases > Infectious diseases: Parasitic > Toxoplasmosis.
· Human Diseases > Infectious diseases: Bacterial > Tuberculosis.
· Human Diseases > Infectious diseases: Viral > Hepatitis C.
· Human Diseases > Infectious diseases: Viral > Measles.
· Human Diseases > Infectious diseases: Viral > Herpes simplex infection.
· Human Diseases > Infectious diseases: Viral > Epstein-Barr virus infection.
· Human Diseases > Cancers: Overview > Pathways in cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Small cell lung cancer. (View pathway)
· Organismal Systems > Development > Osteoclast differentiation. (View pathway)
· Organismal Systems > Immune system > Toll-like receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > NOD-like receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > RIG-I-like receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > IL-17 signaling pathway. (View pathway)
· Organismal Systems > Nervous system > Neurotrophin signaling pathway. (View pathway)
文献引用
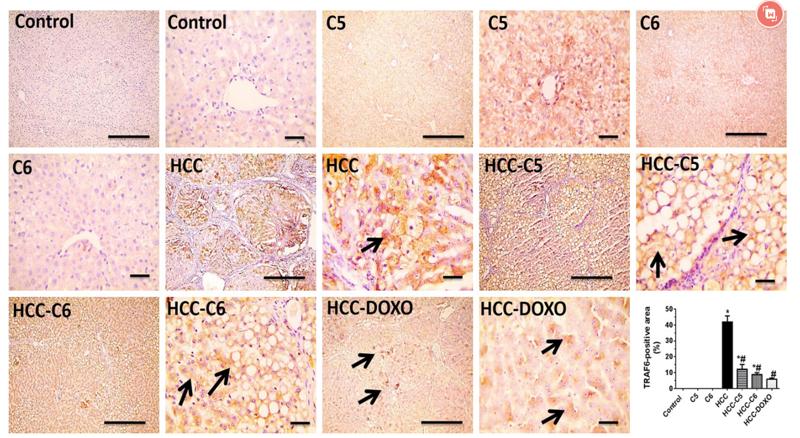
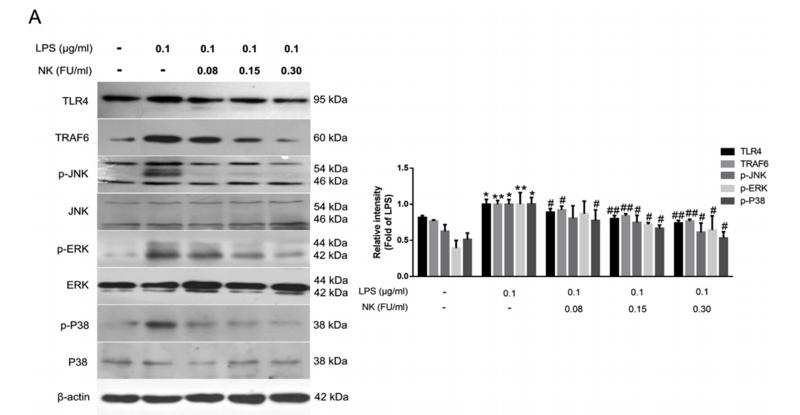
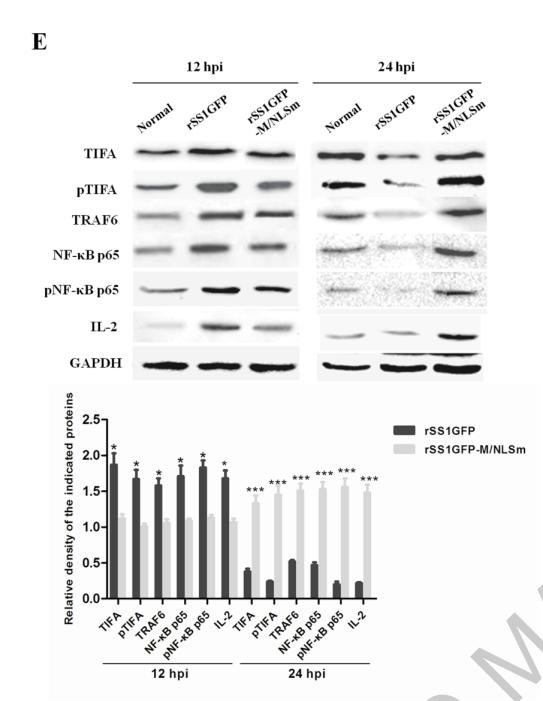
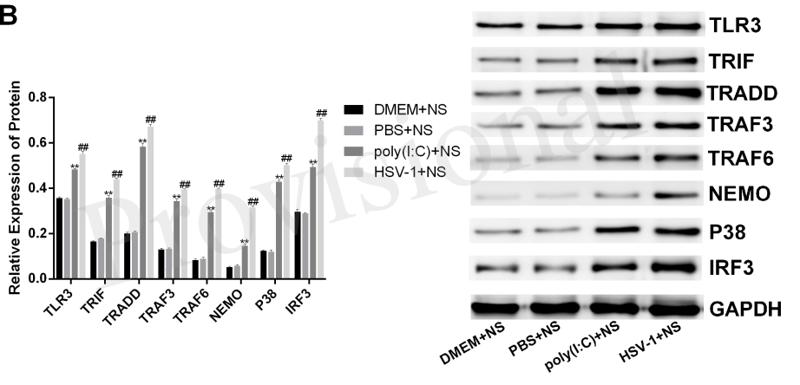
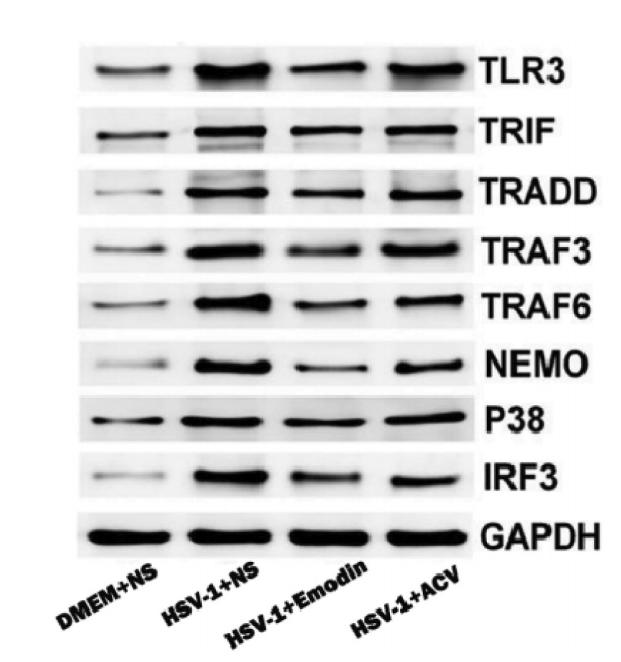
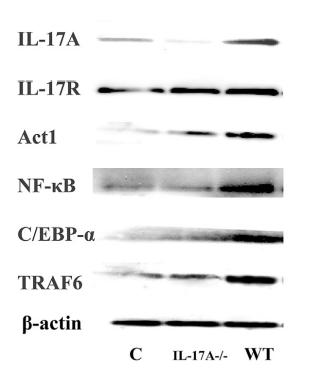
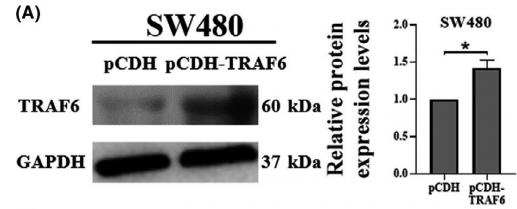
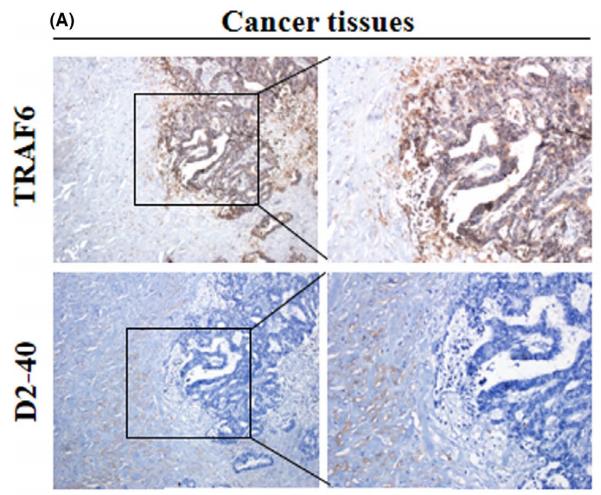
Application: WB Species: Mice Sample: RAW264.7 cells
Application: WB Species: Mice Sample: testes
Application: WB Species: Mouse Sample: BSR-T7/5 cells
限制条款
产品的规格、报价、验证数据请以官网为准,官网链接:www.affbiotech.com | www.affbiotech.cn(简体中文)| www.affbiotech.jp(日本語)产品的数据信息为Affinity所有,未经授权不得收集Affinity官网数据或资料用于商业用途,对抄袭产品数据的行为我们将保留诉诸法律的权利。
产品相关数据会因产品批次、产品检测情况随时调整,如您已订购该产品,请以订购时随货说明书为准,否则请以官网内容为准,官网内容有改动时恕不另行通知。
Affinity保证所销售产品均经过严格质量检测。如您购买的商品在规定时间内出现问题需要售后时,请您在Affinity官方渠道提交售后申请。产品仅供科学研究使用。不用于诊断和治疗。
产品未经授权不得转售。
Affinity Biosciences将不会对在使用我们的产品时可能发生的专利侵权或其他侵权行为负责。Affinity Biosciences, Affinity Biosciences标志和所有其他商标所有权归Affinity Biosciences LTD.